Download: Master Thesis- Martin Tamajka: Segmentation of anatomical organs in medical data
Annotation:
2016, May
Medical image segmentation is an important part of medical practice. Primarily as far as radiologists are concerned it simplifies their everyday tasks and allows them to use their time more effective, because in most cases radiologists only have a certain amount of time they can spend examining patient’s data. Computer aided diagnosis is also a powerful instrument in elimination of possible human failure.
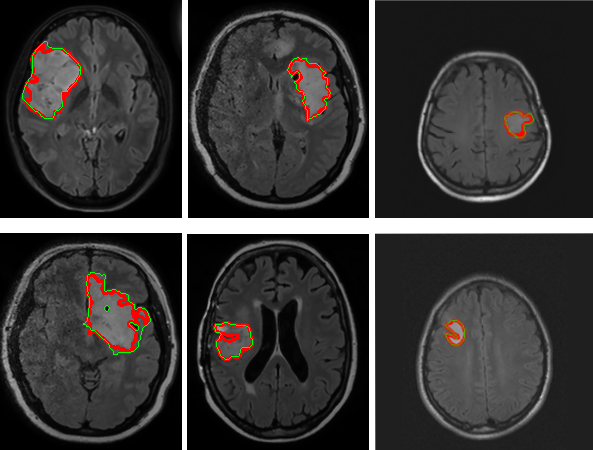
In this work, we propose a novel approach to human organs segmentation. We primarily concentrate on segmentation of human brain from MR volume. Our method is based on oversegmenting 3D volume to supervoxels using SLIC algorithm. Individual supervoxels are described by features based on intensity distribution of contained voxels and on position within the brain. Supervoxels are classified by neural networks which are trained to classify supervoxels to individual tissues. In order to give our method additional precision, we use information about the shape and inner structure of the organ. In general we propose a 6-step segmentation method based on classification.
We compared our results with those of state-of-the-art methods and we can conclude that the results are clearly comparable.
Apart from the global focus of this thesis, our goal is to apply engineering skills and best practices to implement proposed method and necessary tools in such a way that they can be easily extended and maintained in the future.
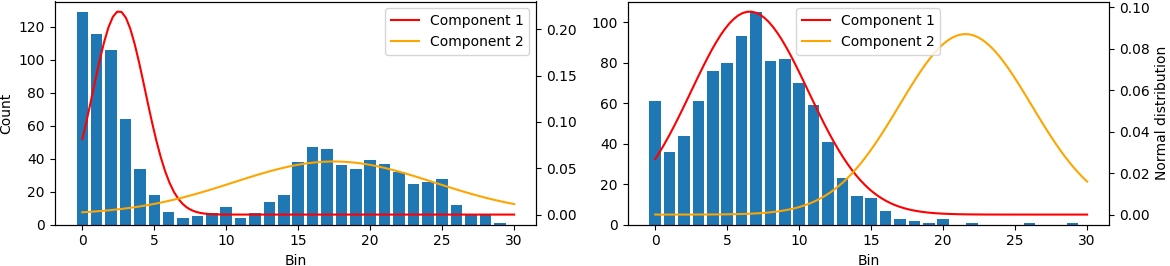 Abstract: The aim of this paper is to propose a novel method to explain, interpret, and support the decision-making process of deep Convolutional Neural Network (CNN). This is achieved by analyzing neuron activations of trained 3D-CNN on selected layers via Gaussian Mixture Model (GMM) … more
Abstract: The aim of this paper is to propose a novel method to explain, interpret, and support the decision-making process of deep Convolutional Neural Network (CNN). This is achieved by analyzing neuron activations of trained 3D-CNN on selected layers via Gaussian Mixture Model (GMM) … more