About
I am currently a finishing Ph.D. student at FIIT STU in Bratislava under the supervision of Dipl. Ing. Vanda Benešová, PhD. I started to incline to computer science and computer vision during my work on my bachelor thesis. In 2015/2016 I was also a member of a team project R3D that cooperated with Austrian research center Joanneum on the topic of 3D Reconstruction and segmentation. The same year’s summer, I attended an Internship at Joanneum Research DIGITAL in Graz where I worked on camera pose estimation and movement models from multiple stereo scans. Later, my master thesis was focused on object segmentation and recognition using RGB-D data. In my dissertation, I focus on more advanced applications of deep learning models in texture similarity and classification and diagnostics of breast cancer. In recent years, I and students under my supervision proposed fine deep learning approaches for breast cancer classification from WSI data, and the application PCD-Quant for quantitative analysis of cilia for Primary Ciliary Dyskinesia, in cooperation with pathologists from Prague histology centers.
Teaching
In the role of Lecturer and Teaching Assistant, I lead the following courses”
- NSIETE – Neural Networks
In the role of Teaching Assistant I teach the following courses:
- PPGSO_B – Principles of Computer Graphics and Image Processing
I used to teach the courses:
- VD_I – Data Visualization
Research
Publications
Marek Jakab, Lukas Hudec, Wanda Benesova, “Partial Disentanglement of Hierarchical Variational Auto-Encoder for Texture Synthesis”, IET Computer Vision, 2021, doi: https://doi.org/10.1049/iet-cvi.2019.0416
Abstract: Multiple research papers have recently demonstrated that deep networks can generate realistically looking textures and stylized images from a single texture example. However, they suffer from some drawbacks. The generative adversarial networks are in general difficult to train. Multiple feature variations encoded in their latent representation require a priori information in order to generate images with specific features. The auto-encoders are prone to generate a blurry output. One of the main reasons is an inability to parametrize complex distributions. We present a novel texture generative model architecture extending the variational auto-encoder approach. It gradually increases the accuracy of details in the reconstructed images. Thanks to the proposed architecture, the model is able to learn a higher level of details resulting from the partial disentanglement of latent variables. The generative model is also capable of synthesizing the complex real-world textures. The model consists of multiple separate latent layers responsible for learning the gradual levels of texture details. Separate training of latent representations increases stability of the learning process and provides partial disentanglement of latent variables. The experiments with proposed architecture demonstrate, the potential of variational auto-encoders in the domain of texture synthesis and also tends to yield sharper reconstruction as well as synthesized texture images.
Jakub Mrva, Stefan Neupauer, Lukas Hudec, Jakub Sevcech and Peter Kapec, “Decision Support in Medical Data Using 3D Decision Tree Visualisation”, The 7th IEEE International Conference on e-Health and Bioengineering EHB 2019, Iasi Romania, 2019, doi: 10.1109/EHB47216.2019.8969926.
Abstract: Decision making is a difficult task, especially in the medical domain. Decision trees are among the most often used supervised machine learning algorithms, primarily due to their straightforward interpretation through rules, they are composed of. Visualizations of decision trees can help with the understanding of data and decision rules on top of them. In this paper we propose a 3D decision tree visualization method that allows effective decision tree exploration and help with decision tree interpretation. The main contribution of this visualization is the ability to display global tree structure and the difference in the distributions of values between subsets created by individual decision nodes. We evaluate our approach using a design study on a typical medical dataset and on a synthetic dataset with controlled properties. We show the visualization’s ability to display individual components of the decision rule and the ability to see other correlated factors in the decision and the change it causes in subsets of data created by individual decision nodes.
D. Hradel, L. Hudec and W. Benesova, “Interpretable Diagnosis of Breast Cancer from Histological Images Using Siamese Neural Networks”, 2019 The 12th International Conference on Machine Vision (ICMV), Amsterdam Netherlands, 2019, doi: 10.1117/12.2557802
Abstract: Breast cancer is one of the most widespread causes of women’s death worldwide. Successful treatment can be achieved only by the early and accurate tumor diagnosis. The main method of tissue diagnosis taken by biopsy is based on the observation of its significant structures. We propose a novel approach of classifying microscopy tissue images into 4 main cancer classes (normal, benign, In Situ and invasive). Our method is based on comparing and determining the similarity of the new tissue sample with previously by specialists annotated examples that are compiled in the collection with other labeled samples. The most probable class is statistically determined by comparing a new sample with several annotated samples. The usual problem of medical datasets is the small number of training images. We have applied suitable dataset augmentation techniques, using the fact that flipping or mirroring of the sample does not change the information about the diagnosis. Our other contribution is that we show the histopathologist the reason why the algorithm has classified tissue into the particular cancer class by ordering the collection of correctly annotated samples by their similarity to the input sample. Histopathologists can focus on searching for the key structures corresponding to the predicted classes.
L. Hudec and W. Benesova, “Texture Similarity Evaluation via Siamese Convolutional Neural Network,” 2018 25th International Conference on Systems, Signals and Image Processing (IWSSIP), Maribor, 2018, pp. 1-5. doi: 10.1109/IWSSIP.2018.8439387
Abstract: Image texture analysis, texture description, and texture similarity evaluation are important areas in computer vision. Evaluation of similarity is needed in many areas as for example object segmentation or image retrieval. We introduce a novel approach for texture similarity measure based on modern deep learning techniques. Our goal is to determine the similarity between patches from homogeneous and also non-homogeneous textures of real-world images. We took the advantage of Siamese Neural Network which is designed to determine the similarity of image pairs. Siamese Neural Network learns to select the most distinctive features responsible for differentiation of the textures. Each of the twin networks creates a feature vector for the image it processes. We used Euclidean and Canberra distance as the similarity metrics to compare these vectors. The final results of the evaluation show a great potential of the proposed method.
URL:Â http://ieeexplore.ieee.org/stamp/stamp.jsp?tp=&arnumber=8439387&isnumber=8439149
Created texture dataset free to download – TexDat
Online sources used to create the dataset
- Free Stock Photos Pexels
- Brodatz Dataset
- SIPI image database
- other sources
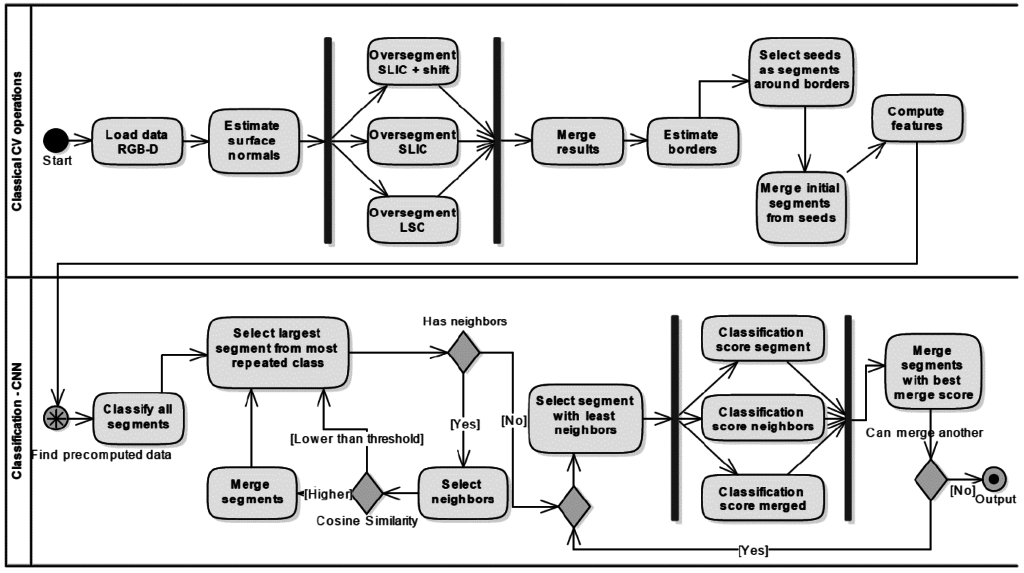
Master Thesis: Automatic segmentation and semantic scene description from RGB-D data
The main goal of this work was to implement unsupervised segmentation and object labeling method. The method specializes in indoor scenes typical for high clutter and overfilling of objects. The proposed approach combines superpixel segmentation of RGB-D images and pseudo-supervised segment merging with help of convolutional neural network. The initial step of merging is based on segments mutual similarity. The next step is iterative semantic merging – jigsaw puzzle approach – two segments are merged only if they are semantically similar – parts of the same object which is evaluated by CNN.
This work was published and presented at IIT.SRC 2017 and presented at SVO? Plze? 2017
Bachelor Thesis: Visual detection of people flow with use of Kinect sensor
This work deals with the task of detecting and tracking people in dense crowd. My solution is based on using depth map (2,5D) acquired by Kinect 1 sensor. Unlike usual approaches, the sensor is positioned above the enter into the area and scanned the space below – bird’s eye view. As for detection, the algorithm searches for human heads in the scan – spatial spherical blobs. After segmentation by morphological reconstruction the found blobs are tracked across video frames using greedy search for most closest blob.